 ,, 蔡霓, 王峰, 农向群
,, 蔡霓, 王峰, 农向群 ,, 王广君, 涂雄兵, 张泽华中国农业科学院植物保护研究所植物病虫害生物学国家重点实验室,北京 100193
,, 王广君, 涂雄兵, 张泽华中国农业科学院植物保护研究所植物病虫害生物学国家重点实验室,北京 100193Expression in vitro of Metarhizium anisopliae Adhesin MAD1 and Its Effect on Inducing Response in Peanut
YAN DuoZi ,, CAI Ni, WANG Feng, NONG XiangQun
,, CAI Ni, WANG Feng, NONG XiangQun ,, WANG GuangJun, TU XiongBing, ZHANG ZeHuaState Key Laboratory for Biology of Plant Diseases and Insect Pests, Institute of Plant Protection, Chinese Academy of Agricultural Sciences, Beijing 100193
,, WANG GuangJun, TU XiongBing, ZHANG ZeHuaState Key Laboratory for Biology of Plant Diseases and Insect Pests, Institute of Plant Protection, Chinese Academy of Agricultural Sciences, Beijing 100193通讯作者:
责任编辑: 岳梅
收稿日期:2020-05-9接受日期:2020-06-10网络出版日期:2021-02-16
| 基金资助: |
Received:2020-05-9Accepted:2020-06-10Online:2021-02-16
作者简介 About authors
闫多子,E-mail:

摘要
关键词:
Abstract
Keywords:
PDF (1016KB)元数据多维度评价相关文章导出EndNote|Ris|Bibtex收藏本文
本文引用格式
闫多子, 蔡霓, 王峰, 农向群, 王广君, 涂雄兵, 张泽华. 绿僵菌黏附素MAD1体外表达及诱导花生响应的作用[J]. 中国农业科学, 2021, 54(4): 744-753 doi:10.3864/j.issn.0578-1752.2021.04.007
YAN DuoZi, CAI Ni, WANG Feng, NONG XiangQun, WANG GuangJun, TU XiongBing, ZHANG ZeHua.
开放科学(资源服务)标识码(OSID):

0 引言
【研究意义】绿僵菌经过漫长的演化过程,形成了兼备腐生、植物共生、昆虫致病的生活能力,以适应各种环境[1]。由于具有广泛的昆虫寄主和高效致病力,已被研究用于防治几十种农林害虫[2],并作为昆虫病原真菌的模式菌用于研究对昆虫的侵染过程和致病机理。MAD1作为绿僵菌黏附及毒力的调节蛋白,参与绿僵菌与昆虫寄主的互作,但绿僵菌与植物的互作机制极少报道。研究MAD1诱导植物响应对调控绿僵菌与植物的互作具有重要意义。【前人研究进展】绿僵菌侵染昆虫寄主首先是在表皮上附着、定殖,然后穿透表皮进入体内[3]。已有研究表明,绿僵菌在附着昆虫体表时产生黏附蛋白MAD1,促进真菌孢子稳定地吸附,有助于形成的附着胞“钉”入宿主体表,对成功侵入寄主起关键作用。当缺失mad1后,绿僵菌分生孢子对寄主体表的黏附能力与毒力均显著降低[4,5,6]。用体外表达的MAD1蛋白与麦麸做成饵剂饲喂蝗虫,处理后24 h检测发现,蝗虫中肠的过氧化物酶(peroxidase,POD)、超氧化物歧化酶(superoxide dismutase,SOD)活性受到抑制,表明MAD1对蝗虫的免疫有一定的抑制作用[7]。近年研究发现,绿僵菌能够在植物根际和进入植物体内宿存,且促进植物生长。例如,用绿僵菌分生孢子处理柳枝稷(Panicum virgatum)和菜豆(Phaseolus vulgaris)后,可促进根毛及其侧根生长[8]。绿僵菌孢子悬浮液处理番茄(Lycopersicon esculentum)种子后,种植的番茄株高、根长、侧根数量以及根干重均比对照明显增加[9]。另外,绿僵菌侵入植物根组织内的现象也得到证实。以绿色荧光蛋白(green fluorescent protein,GFP)标记的罗伯茨绿僵菌(Metarhizium robertsii)菌株接种菜豆幼苗60 d后,用共聚焦显微镜观察到根表皮的绿色荧光菌丝;将菜豆根系组织制成切片,用溴酚蓝染色法观察到菌体在60 d内穿透豆根细胞壁,侵入根细胞内生长[8]。以增强型绿色荧光蛋白(enhanced green fluorescent protein,eGFP)标记的金龟子绿僵菌(Metarhizium anisopliae)接种玉米(Zea mays)14 d后,通过定性和定量PCR从玉米根DNA中扩增到egfp片段,通过激光共聚焦(laser scanning confocal microscopy,LSCM)技术检测到定殖于玉米根中带绿色荧光的菌丝,证明绿僵菌可以定殖于玉米根部[10]。然而,有关绿僵菌作为植物内生菌如何黏附、定殖于植物的过程和机理研究甚少,植物对绿僵菌诱导产生的分子生物学响应也极少报道。植物经长期进化形成了应对微生物的免疫系统。位于细胞膜上的各类模式识别受体(pattern-recognition receptor,PRR)能够感知微生物相关分子模式(microbe-associated molecular pattern,MAMP),激发病原因子触发免疫反应(PAMP-triggered immunity,PTI),如活性氧(reactive oxygen species,ROS)的爆发、钙离子(Ca2+)的流入、丝裂原活化蛋白激酶(mitogen-activated protein kinase,MAPK)的激活、防御基因表达的上调等[11]。微生物为了成功入侵宿主,又进化出效应蛋白来躲避或抑制植物免疫反应,以在植物体内宿存和生长。而植物也进化出相应的抗病蛋白(R蛋白)来应对效应子,激活效应子触发免疫反应(effector-triggered immunity,ETI),抵抗微生物入侵[12]。对于致病或有益的微生物,植物可启动或保留一定的反应来抵抗或兼容[13,14]。有研究表明有益真菌木霉(Trichoderma sp.)在定殖拟南芥(Arabidopsis thaliana)的过程中,是通过对植物免疫反应的短暂抑制来实现在根部的定殖[15]。根瘤菌会主动抑制宿主的免疫防御反应,使感染和共生得以建立[16]。【本研究切入点】前期对绿僵菌处理的花生(Arachis hypogaea)根组织进行转录组测序分析,发现花生的一些代谢途径和免疫途径基因出现显著差异表达[17]。以绿僵菌可定殖植物为出发点,研究黏附素MAD1在介导绿僵菌黏附、定殖中作为分子信号诱导植物产生的分子生物学响应。【拟解决的关键问题】克隆mad1并进行体外真核表达,检测花生免疫防御等相关基因对表达蛋白诱导的响应,明确MAD1蛋白在绿僵菌定殖植物过程中的作用,为阐述绿僵菌与植物共生关系建立的机制提供基础数据。1 材料与方法
试验于2018—2019年在中国农业科学院植物保护研究所植物病虫害生物学国家重点实验室完成。1.1 生物材料
金龟子绿僵菌IPPM010202菌株由本实验室分离鉴定,保存于马铃薯蔗糖琼脂培养基(PSAY)中。PSAY为1 L水中含马铃薯200 g,蔗糖20 g,琼脂18 g,酵母粉5 g。产菌丝培养基为1 L水中含MgSO4 2 g,蔗糖20 g,酵母粉10 g,K2HPO4 5 g。酵母培养基YPD为1 L水中含酵母提取物10 g,蛋白胨20 g,葡萄糖20 g。酵母表达培养基BMGY为1 L水中含酵母粉10 g,KH2PO4 11.8 g,蛋白胨20 g,甘油10 mL。酵母表达培养基BMMY为1 L水中含酵母粉10 g,KH2PO4 11.8 g,蛋白胨20 g,甲醇5 mL。基础葡萄糖培养基MD为1 L水中含酵母粉13.4 g,生物素0.4 mg,葡萄糖20 g。
大肠杆菌感受态DH5α购自全式金生物技术有限公司。PMD19-T载体、Prime ScriptTM 1st strand cDNA sythesis Kit反转录试剂盒、限制性内切酶EcoR I、Not I、Sac I、T4 DNA连接酶购自TaKaRa公司。酵母表达载体pPIC9K、毕赤酵母宿主菌GS115购自苏州泓迅生物科技有限公司。D-山梨醇、Biomed回收试剂盒购自北京威莱博生物技术有限公司。
供试花生为鲁花11号品系,从农资商店采购。
1.2 菌丝培养及总RNA提取
将绿僵菌孢子接种到产菌丝培养基中,28℃ 160 r/min摇床培养60 h。培养液通过抽滤尽量去除水分,然后用TRIzol?分离试剂盒(Invitrogen)提取总RNA。用Prime ScriptTM 1st strand cDNA sythesis Kit反转录试剂盒(TaKaRa公司)进行反转录,得到cDNA,并用NanoPhotometer微量分光光度计(IMPLEN,德国)检测浓度。1.3 黏附素基因mad1的克隆
以绿僵菌转录组的Unigenes作为参考序列[17],在GenBank中进行同源性检索比对,找到mad1的开放阅读框(ORF),设计扩增全长特异性引物mad1-F(5′-TG CCGAGAGGTGCTCAGCTATTC)和mad1-R(5′-TAGC ACTGTTGTTTAACGGCCGAAG)。优化后的PCR反应体系(50 μL):2×Taq PCR Master Mix 25 μL,mad1-F 4 μL,mad1-R 4 μL,cDNA 2 μL,补足ddH2O 至50 μL。PCR反应的温度控制程序为94℃预变性5 min,随后,94℃变性30 s,55℃退火30 s,72℃延伸2 min,35个循环,72℃延伸10 min。PCR产物经1.5%琼脂糖凝胶电泳,用Biomed回收试剂盒进行纯化回收。再连接到PMD19-T载体上,热激转化导入大肠杆菌感受态DH5α中。通过氨苄(Amp)抗性筛选阳性克隆,PCR检验后送至上海生工生物(北京)公司测序。1.4 生物信息学分析
以DNAMAN对克隆得到的mad1序列进行蛋白翻译。使用在线工具ProtParam(http://web.expasy.org/ protparam)预测蛋白氨基酸组成、分子量、理论等电点、不稳定指数和总平均亲水性等性质;利用SMART(http://smart.embl-heidelberg.de/)在线程序分析其结构域。1.5 黏附素基因mad1的真核表达
1.5.1 真核表达载体构建与酵母转化 利用表达载体pPIC9K中AOX1作为启动子,在克隆的mad1 5′端增加限制性内切酶位点EcoR I,在3′端增加his-tag、终止密码子TGA和限制性内切酶位点Not I,得到EcoR I-mad1-Not I序列,将EcoR I-mad1-Not I与载体pPIC9K在限制性内切酶EcoR I和Not I作用下分别进行双酶切,酶切后产物于16℃条件下过夜连接。连接产物热激转化导入大肠杆菌感受态DH5α中。通过Amp抗性筛选阳性转化子送至上海生工生物(北京)公司测序,确定mad1位置、方向、大小和读码框完全正确。用质粒DNA小量提取试剂盒(天漠生物)提取质粒,pPIC9K-mad1重组表达质粒用Sac I内切酶线性化,用Biomed回收试剂盒进行纯化回收。将线性化的质粒通过电击转化法转入毕赤酵母GS115感受态细胞中。涂布于MD平板,48 h后观察并挑选菌落。1.5.2 转化子筛选 挑取MD平板上5个单克隆,接种到YPD培养基中,28℃培养36 h。离心收集菌体,用TIANamp Yeast DNA Kit酵母基因组DNA提取试剂盒(天根公司)提取酵母基因组。从重组质粒pPIC9K-mad1上mad1的5′端及接头序列设计特异性引物cmad1-F(5′-ATGAAGTCTGCTCTTTCTGTTGT TGTTGCCGC-3′),以mad1的3′端及外延15 bp设计反向引物cmad1-R(5′-GACCGGTCTTCTCGTGCGG CCGCTCAGTGGTGGTGGTGGTGGTG-3′),进行PCR扩增验证。PCR反应体系为2×Taq PCR Master Mix 25 μL,cmad1-F 4 μL,cmad1-R 4 μL,cDNA 2 μL,补足ddH2O至50 μL。温度控制程序为94℃预变性5 min,随后,94℃变性30 s,55℃退火30 s,72℃延伸2 min,35个循环,72℃延伸10 min。产物用1.5%琼脂糖电泳,鉴定阳性克隆。
1.6 重组蛋白MAD1的诱导表达和纯化
阳性克隆单菌落接种到含有100 mL BMGY培养基的锥形瓶中,于28℃ 250 r/min培养16—20 h。8 000 r/min离心收集菌体,加1 mL BMMY培养基重悬,转接到100 mL BMMY培养基继续培养,并间隔24 h向培养基中添加甲醇至终浓度1.0%,诱导培养72 h。培养物经离心分离,分别收集上清和沉淀,用10% SDS-PAGE检测。沉淀的菌体用缓冲液(20 mmol·L-1 Tris,500 mmol·L-1 NaCl,pH 8.0)重悬,进行超声波破碎。破碎后的菌体4℃下离心,收集上清液,用Ni-NTA镍柱纯化,得到MAD1蛋白,保存在缓冲液(20 mmol·L-1 Tris,500 mmol·L-1 NaCl,pH 7.8)中备用。
1.7 表达蛋白诱导花生响应的作用分析
1.7.1 MAD1诱导处理 花生种植参照MICHé等[18]的方法先用4% NaClO和15% H2O2溶液进行种子表面消毒,再将种子浸泡于含50 mL蒸馏水的15 cm×15 cm的培养皿中,黑暗培养箱中28℃催芽24 h,然后挑选萌芽健康一致的种子,播种到装有已灭菌蛭石的花盆中,于培养室(L﹕D=14﹕10,28℃)培养14 d,相对湿度为40%—60%,生长至二叶期。MAD1处理花生:将纯化的MAD1蛋白用50 kD蛋白超滤管(Millipore)离心浓缩,更换缓冲液为ddH2O,配置成20 μg·mL-1的处理液。取上述生长一致的花生幼苗6株,每2株一组,将根部浸入20 mL的蛋白液中,3个重复处理,同时以ddH2O处理为对照。于28℃处理0.5、6、12、24 h,取出幼根,用吸水纸吸干,根部剪成5—8 mm,立即置于液氮中。同一处理不同的重复进行混样后,分成3份,保存于-80℃备用。
1.7.2 花生根实时荧光定量PCR检测 取花生根样品,置于液氮中研磨,然后用TRIzol?分离试剂盒(Invitrogen)提取总RNA。用Prime ScriptTM 1st strand cDNA sythesis Kit反转录试剂盒(TaKaRa公司)进行反转录,得到cDNA。并用NanoPhotometer微量分光光度计(IMPLEN,德国)检测浓度。
采用实时荧光定量PCR(qPCR)检测不同处理的花生根组织中免疫防御相关基因转录水平。基于之前的转录组分析[17]确定候选基因,设计的引物序列见表1,内参基因用60s ribosomal protein L7(RPL7)。用SYBR Green荧光定量试剂盒(TaKaRa公司)在7500型(ABI公司)实时荧光定量PCR系统进行反应。反应体系为SYBR Green preMix 10 μL,正反向引物各2 μL,cDNA 1 μL,补足ddH2O至20 μL。PCR扩增程序为95℃预变性3 min,95℃变性15 s,60℃复性1 min,40个循环。每样品3次技术重复,基因相对表达量采用2-ΔΔCt[19]的方法进行分析。
Table 1
表1
表1候选基因及其特异性引物
Table 1
| 基因Gene | 引物序列Primer sequence | 基因Gene | 引物序列Primer sequence | |
|---|---|---|---|---|
| CERK1 | 5′-CTACTCAGCCATTGCCAGTG | RPK | 5′-AAGGTGGAGTTTGTTATGCC | |
| 5′-CAGTCCTAAGAGGGTATGTGATGA | 5′-CCTAAGGACTTGAGACCCTG | |||
| PTi1 | 5′-TTTGACCTGGGCACAGAGAG | RML1A | 5′-CAGTGAGCGTTGCGAGATAC | |
| 5′-GACACGGGTGGAATGAAGAC | 5′-TTGGGTTTCTTTCAGCGACT | |||
| SCW1 | 5′-GAGTAAGCGATTGCGGACAG | CDPK | 5′-CAAGACCATTGTTGAGGTTG | |
| 5′-TGTTCTCTCCCAAGTTCCCA | 5′-CTCCACACATCTGATTCTGG | |||
| MMK1 | 5′-AACTCGGAGACGAATGAGCA | MAPK | 5′-CTTCCACAATGTCCAAAGCA | |
| 5′-TTGGTGAAGGTCAGTGTCCA | 5′-CCAGATGAGTTCCTTGATGTC | |||
| MYB86 | 5′-CCACAACAGACAACAGTTTACG | RPL7 | 5′-AGAAGAGGGAGGAGGAATGG | |
| 5′-TAGAGCACCATCTCCTACCTC | 5′-GGATGCGGATGATAAACAGG |
新窗口打开|下载CSV
qPCR检测每个样品的3次技术重复。以Student’s t-test(P0.05)(SPSS version 16.0)方法分析数据的统计学显著性差异,所有试验相同处理不同重复的数据采用均值±标准误(standard error,SE)表示。
2 结果
2.1 黏附素基因mad1的克隆和生物信息学分析
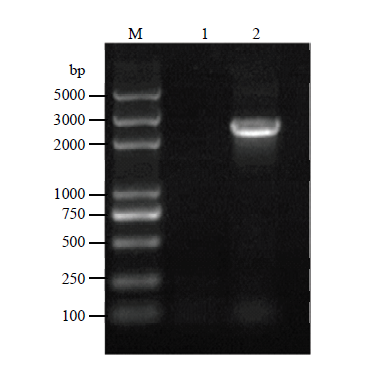
用所设计的mad1特异性引物,从金龟子绿僵菌IPPM010202菌株cDNA扩增得到了预期长度的扩增片段(图1)。经测序,该片段包含mad1的完整开放阅读框(ORF),全长为2 136 bp(GenBank序列号MT375602)。该基因编码711个氨基酸,该蛋白的理论分子量为74.8 kD,预测等电点pI为6.28。在20种氨基酸中,苏氨酸(Thr)所占比例最高(12.0%),甲硫氨酸(Met)和色氨酸(Trp)所占的比例最低(0.7%);含51个负电荷氨基酸残基,48个正电荷氨基酸残基;不稳定指数为44.66,属于不稳定蛋白;总平均亲水性-0.414,属于亲水性蛋白。经二级结构分析,MAD1在N端有信号肽,在C端有预测的糖基磷脂酰肌醇(GPI)细胞壁锚定位点,在中间区域包含苏氨酸(Thr)的串联重复序列。在末端区域含脯氨酸(Pro)串联重复序列。另外,MAD1结构中有一个含有8个半胱氨酸的CFEM功能结构域,含这种结构域的蛋白可以作为细胞表面受体、信号转导者或与宿主相互作用的黏附分子(图2)。图1
 新窗口打开|下载原图ZIP|生成PPT
新窗口打开|下载原图ZIP|生成PPT图1绿僵菌mad1基因全长扩增
Fig. 1Amplification of the mad1 full-length gene from M. anisopliae
M:DL5000 DNA marker;1:ddH2O;2:绿僵菌cDNA cDNA from M. anisopliae
图2
 新窗口打开|下载原图ZIP|生成PPT
新窗口打开|下载原图ZIP|生成PPT图2绿僵菌黏附素MAD1结构
Fig. 2Structure of M. anisopliae adhesin MAD1
2.2 重组表达质粒和酵母重组子的鉴定
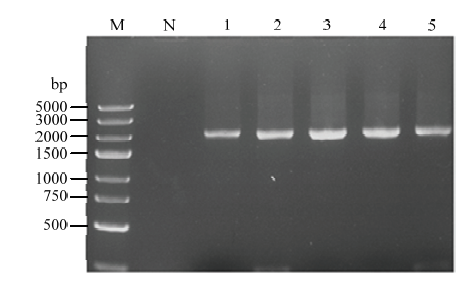
将mad1构建到表达载体pPIC9K上,获得了重组质粒,经EcoR I和Not I双酶切,电泳检测到预期大小一致的片段。将阳性重组质粒送检测序,结果证明mad1的插入位置、方向、大小和读码框均正确,表明重组表达载体构建成功。转入宿主毕赤酵母GS115后,挑选单克隆,以cmad1-F、cmad1-R为引物进行PCR检测,结果均在预期的2 136 bp附近得到单一条带(图3),表明已将重组质粒转入宿主毕赤酵母中。图3
 新窗口打开|下载原图ZIP|生成PPT
新窗口打开|下载原图ZIP|生成PPT图3mad1转化酵母阳性克隆PCR检验
Fig. 3Examination of positive clones of mad1 transformed yeast by PCR
M:DL 5000 DNA marker;N:空载酵母GS115 Empty carrier yeast GS115;1—5:重组单克隆Recombinant clones
2.3 诱导表达产物的检验
含有mad1的重组酵母菌株经甲醇诱导表达,分别对发酵液和菌体蛋白进行SDS-PAGE分析,结果显示,发酵液上清中未检测到目的蛋白(图4-A)。发酵液沉淀的菌体经超声波破碎后,上清中检测到目的蛋白(图4-B),经镍柱纯化后SDS-PAGE检测,在20 mmol·L-1咪唑洗脱液中获得大量蛋白,符合预期分子量74.8 kD(图4-C),表明mad1在毕赤酵母中成功表达。图4
 新窗口打开|下载原图ZIP|生成PPT
新窗口打开|下载原图ZIP|生成PPT图4重组酵母诱导表达MAD1的SDS-PAGE分析
Fig. 4SDS-PAGE analysis of recombinant yeast induced expression of MAD1
A:发酵液上清Supernatant of fermentation broth;B:菌体破碎提取上清Extract of crushed cell;C:MAD1蛋白纯化Purification of MAD1 protein
2.4 黏附素MAD1诱导花生响应的作用分析
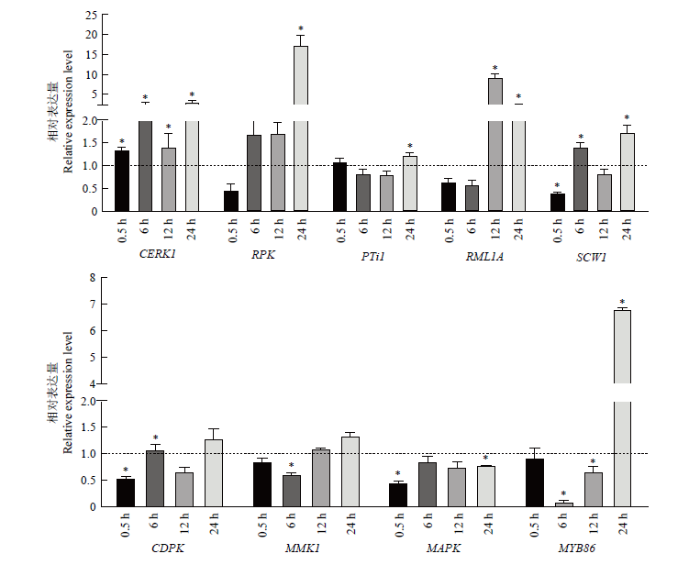
以重组酵母表达的MAD1处理花生幼苗根部,检测了在处理0.5、6、12、24 h的4个时间段内花生根组织9个免疫防御相关基因的相对表达量变化。结果如图5所示,在与MAD1互作初期的0.5 h内,花生根膜受体基因CERK1已经启动,表达量显著高于对照,至6 h时继续上升至对照的2.5倍,之后有所波动,一直保持显著高于对照的水平。另一膜受体基因RPK的表达是0.5 h内先略受抑制,然后迅速上升,6—12 h时达到对照的1.69倍左右,24 h时陡增至17倍以上,呈现强烈反应。对于防御相关基因PTi1、RML1A和SCW1,初始时基本无反应或受到抑制,PTi1受抑制时间较长,直至24 h才出现小幅上调表达;RML1A在12 h时凸显9倍的上调峰值,24 h仍有2倍的上调表达;SCW1的表达量在下降、上升间转换。对于信号转导相关的CDPK、MMK1和MAPK,互作初期0.5 h时,CDPK和MAPK出现下调表达,MMK1基本无变化;随后,CDPK为0.54—1.29倍幅度的波动,MMK1在6 h下调表达0.62倍后转为逐步上升至1.34倍,MAPK的表达量有回升但始终未超过对照。转录因子基因MYB86在初始0.5 h时无反应,但随后出现先下降后上升的剧烈反应,6 h时表达量下调至对照的0.1倍,12 h时回升至0.66倍,再快速提升,24 h时达到6.8倍的上调表达。由此可见,MAD1与花生互作的0.5—24 h内,已经诱导了花生根组织免疫相关基因的回应,反映在膜受体基因CERK1、RPK转录水平的快速和显著大幅的上调,以及防御相关蛋白基因PTi1、RML1A、SCW1和转录因子基因MYB86相应的调节。图5
 新窗口打开|下载原图ZIP|生成PPT
新窗口打开|下载原图ZIP|生成PPT图5MAD1诱导花生相关基因的相对表达量
对照的表达量为1,虚线表示;“*”表示相对于对照有显著差异,P<0.05
Fig. 5The relative expression of related genes induced by MAD1
The expression level of the control is 1, indicated by the dotted line. Symbol ‘*’ indicates a significant difference from the control, P<0.05
3 讨论
微生物与植物的互作始于化学信号之间的识别和交换。本研究表明,绿僵菌黏附素MAD1是绿僵菌与花生互作初始的一个重要信号。然而,目前有关绿僵菌与植物互作机理知之甚少,对黏附素MAD1在互作中的作用未见报道。本研究克隆并分析了MAD1结构,发现在N端含有疏水信号肽(SP)序列,引导蛋白质的分泌及跨膜转移。同时C端有高度保守的糖基磷脂酰肌醇(GPI)锚定位点。研究报道GPI锚定蛋白通常被磷酸化和糖基化,蛋白外壳由β-1,6-葡聚糖、β-1,3-葡聚糖和少量甲壳素构成的多聚糖层包围,且蛋白外壳通过GPI锚以共价键的方式与β-1,6糖苷键相连,同时依次与β-1,3-葡聚糖的分子末端相连,将蛋白锚定在细胞膜上,在真菌对宿主的黏附、侵袭和自身形态转换中起重要作用[20,21]。MAD1结构中的CFEM功能结构域,含有8个半胱氨酸,是真菌细胞膜蛋白特有结构。作为细胞表面受体或信号转导器,参与细胞的黏附作用[22,23]。一些含有CFEM的真菌蛋白在侵染植物中起重要作用,例如稻瘟病菌Pth11蛋白,含有CFEM结构域,是侵染植物过程中附着胞发育所必需的[24],这些结构预测为进一步研究MAD1的功能提供了依据。植物通过膜表面识别受体感知和识别外来微生物化学信号,启动下游信号转导通路,激活或抑制相关酶及蛋白活性,协调共生与防御的关系。膜表面模式识别受体(PRR)有多种类型,包括CERK1、RPK,不同的PRR已被证明可以激活细胞免疫信号通路。CERK1(几丁质激发子受体激酶)具有LysM(赖氨酸基序),有跨膜激活作用,通过与CEBiP(几丁质激发子结合蛋白)或LYP4/6形成二聚体,来识别微生物表面的几丁质和肽聚糖类物质,引起活性氧爆发和触发MAPK级联信号转导[25,26,27]。RPK含有亮氨酸重复结构域,位于细胞膜表面感知外界信号,可作为ABA信号的调节因子。RPK1功能丧失,植物在种子萌发、植株生长、气孔关闭和基因表达方面存在ABA敏感性[28]。而ABA通路由4个核心组分共同组成一个双重负调控系统,调节ABA对非生物胁迫包括机械压力、损伤等耐受性的应答[29]。本研究发现,在绿僵菌MAD1处理的花生根组织中,首先是CERK1表现出感应识别,在0.5 h时已经显著上调表达,而RPK的表达在0.5 h内先略受抑制,然后迅速上升,24 h时强烈上调至17倍以上。这两种膜受体上调表达引发了下游相应的信号转导,从而协调防御和共生反应。
植物免疫系统在植物与微生物的互作中起着核心作用。在识别微生物相关分子模式(MAMP)时,植物会触发免疫应答(MTI)。对于致病或有益的微生物,膜表面受体通过识别特定信号分子,可启动或保留一定的反应来抵抗或兼容[14]。植物通过PRR对MAMP识别后,招募和调节受体激酶复合物,来激活多层免疫信号级联反应。MAPK和MMK1可以磷酸化并激活促丝裂原活化蛋白,级联控制了许多下游免疫信号事件,是许多微生物效应的靶标[30]。CDPK是Ca2+依赖蛋白激酶,涉及钙信号转导途径。CDPK与MAPK可以共同调控下游转录因子及植物早期防御反应基因的表达。尤其是胼胝质沉积、抗菌化合物积累和防御激素的调节等[31,32,33]。有研究表明,MAPK级联效应的阻断可以抑制下游免疫基因的激活和相关防御基因的表达。例如大豆MPK4调控WRKY、MYB和bHLH转录因子的表达,进而抑制防御相关基因的表达[34]。还有研究表明许多来自有益微生物的共生分子可以抑制MAMP引发的根免疫反应。如根瘤菌对植物细胞的感染是由根瘤菌多糖、鞭毛蛋白和真菌甲壳素、NopL蛋白(结节蛋白L)化合物引起的,这些化合物能够抑制宿主植物的防御反应[35]。日本短根瘤菌(Bradyrhizobium japonicum)的Nod因子可以强烈抑制大豆(Glycine max)和拟南芥中各种MAMP诱导的免疫反应[36]。根瘤菌的共生分子(LPS)不仅抑制植物早期活性氧的爆发,而且还抑制了后期防御相关基因转录重编程[37]。本研究发现在MAD1作用的24 h内,信号转导相关基因MAPK、CDPK和MMK1在初始时都先下调表达,然后有所回升,处于对照表达水平之下或小幅上下波动。表明在MAD1处理的初始0.5—12 h,花生的免疫反应被短暂的抑制,推测MAD1可能作为一种共生因子参与真菌在植物中的定殖。MYB86是一种转录阻遏子,应答环境变化与激素调节,与植物防卫反应有关。PTi1、RML1A是防御相关蛋白。PTi1可与Pro互作参与抗病反应,RML1A是NBS-LRR类型的抗病蛋白,该类蛋白参与植物天然免疫系统的效应触发免疫(ETI),SCW1是膜整合蛋白,与细胞壁的加固和完整性有关,涉及到植物细胞壁的破坏及重建。本研究中,在MAD1抑制信号转导的同时,下游转录因子基因MYB86、抗病蛋白基因PTi1和RML1A在0.5—6 h均出现显著下调表达,而RML1A和 SCW1分别在12 h和24 h后出现急剧上调,说明MAD1作用初期诱导了花生免疫抑制因子参与兼容绿僵菌的定殖,且植物防御相关基因在短时间内下调,而随后激活了效应触发免疫,增强植物抗病性,并可能通过SCW1启动了共生建立过程所需的细胞壁修复与重建。事实上,绿僵菌在植物上的定殖和建立共生是一个动态的相互作用过程,涉及营养、防御等多方面信号交流和响应,期待今后有更进一步的研究阐释。
4 结论
成功克隆mad1并构建了绿僵菌MAD1真核表达系统,表达并纯化出具有生物活性的MAD1蛋白。通过处理花生根证明了MAD1具有激活花生根膜受体基因CERK1和RPK的转录响应,早期抑制免疫相关基因MAPK、CDPK、MMK1及防御相关基因如PTi1、RML1A的转录水平,有利于绿僵菌在花生根组织上定殖;另外MAD1可调控细胞壁整合基因SCW1表达,参与根上受损细胞壁的修复与重建,促进绿僵菌与花生共生关系的建立。诱导植物的免疫抑制和细胞壁重建整合可能是绿僵菌与植物建立共生关系的重要步骤。参考文献 原文顺序
文献年度倒序
文中引用次数倒序
被引期刊影响因子
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 2]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 2]
[本文引用: 1]
[本文引用: 1]
[本文引用: 3]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
[本文引用: 1]
